 |
 |
Have we entered the era of the 1 year sequencer release cycle? *Updated*
Tuesday, January 13, 2015
Illumina's $1000 Genome*
Wednesday, January 15, 2014
A coming of age for PacBio and long read sequencing? #AGBT13
Saturday, February 23, 2013
Next Generation Sequencing rapidly moves from the bench to the bedside #AGBT13
Friday, February 22, 2013
#AGBT day one talks and observations: WES/WGS, kissing snails, Poo bacteria sequencing
Wednesday, February 20, 2013
Got fetal DNA on the brain?
Friday, September 28, 2012
Memes about 'junk DNA' miss the mark on paradigm shifting science
Friday, September 7, 2012
So, you've dropped a cryovial or lost a sample box in your liquid nitrogen container...now what?
Thursday, August 16, 2012
A peril of "Open" science: Premature reporting on the death of #ArsenicLife
Thursday, February 2, 2012
Antineoplastons? You gotta be kidding me!
Thursday, October 27, 2011
YouTube: Just a (PhD) Dream
Thursday, October 27, 2011
Slides - From the Bench to the Blogosphere: Why every lab should be writing a science blog
Wednesday, October 19, 2011
Fact Checking AARP: Why soundbytes about shrimp on treadmills and pickle technology are misleading
Monday, October 17, 2011
MHV68: Mouse herpes, not mouth herpes, but just as important
Monday, October 17, 2011
@DonorsChoose update: Pictures of the materials we bought being used!!
Friday, October 14, 2011
Is this supposed to be a feature, @NPGnews ?
Tuesday, October 4, 2011
A dose of batshit crazy: Bachmann would drill in the everglades if elected president
Monday, August 29, 2011
A true day in lab
Wednesday, August 10, 2011
A day in the lab...
Monday, August 8, 2011
University of Iowa holds Science Writing Symposium
Tuesday, April 26, 2011
Sonication success??
Monday, April 18, 2011
Circle of life
Thursday, March 17, 2011
Curing a plague: Cryptocaryon irritans
Wednesday, March 9, 2011
Video: First new fish in 6 months!!
Wednesday, March 2, 2011
The first step is the most important
Thursday, December 30, 2010
Have we really found a stem cell cure for HIV?
Wednesday, December 15, 2010
This paper saved my graduate career
Tuesday, December 14, 2010
Valium or Sex: How do you like your science promotion
Tuesday, November 23, 2010
A wedding pic.
Tuesday, November 16, 2010
To rule by terror
Tuesday, November 9, 2010
Summary Feed: What I would be doing if I wasn't doing science
Wednesday, October 6, 2010
"You have more Hobbies than anyone I know"
Tuesday, October 5, 2010
Hiccupping Hubris
Wednesday, September 22, 2010
A death in the family :(
Monday, September 20, 2010
The new lab fish!
Friday, September 10, 2010
What I wish I knew...Before applying to graduate school
Tuesday, September 7, 2010
Stopping viruses by targeting human proteins
Tuesday, September 7, 2010
 |
 |
 |
 |
Brian Krueger, PhD
Columbia University Medical Center
New York NY USA
Brian Krueger is the owner, creator and coder of LabSpaces by night and Next Generation Sequencer by day. He is currently the Director of Genomic Analysis and Technical Operations for the Institute for Genomic Medicine at Columbia University Medical Center. In his blog you will find articles about technology, molecular biology, and editorial comments on the current state of science on the internet.
My posts are presented as opinion and commentary and do not represent the views of LabSpaces Productions, LLC, my employer, or my educational institution.
Please wait while my tweets load 
 |
 |
youtube sequencing genetics technology conference wedding pictures not science contest science promotion outreach internet cheerleaders rock stars lab science tips and tricks chip-seq science politics herpesviruses
 |
 |
 |
 |
How AAAS and Science magazine really feel about sexual harassment cases in science
In 1990, the scientific community embarked on a landmark experiment to completely sequence the human genome. At the time, it was assumed that knowing the exact sequence of the human genome would provide scientists all of the information they ever wanted to know about genomics and how DNA contributes to human disease. At least this is how the project was presented to the public, however, every genome scientist knew that obtaining the sequence of the human genome was a lot like getting a cake recipe that listed all of the ingredients but didn't explain at all how much of each ingredient to use, how to mix the ingredients together or how long to bake that mixture to create a delicious cake. Compound that with the fact that the list contains over 35,000 ingredients and you can quickly understand why the sequence alone wasn’t very informative. We spent over 3 billion dollars to obtain this sequence and since the final publication of the sequence in 2003, the media has questioned its usefulness. Apparently, because cancer wasn’t cured in 10 years, the human genome project is largely seen by the media as a colossal waste of money; however, obtaining this sequence has been invaluable in speeding up research and has significantly contributed to our understanding of how some of these ingredients function to affect human disease.
One of the important new themes in the decade since the completion of the human genome is that the actual sequence of the DNA is even less important than we ever imagined. I say this largely because the 35,000 or so genes coded by the DNA represent only 1-2% of the actual code. That means that 98-99% of our DNA is either useless Junk DNA (not very likely), or it contributes to life in ways that we yet don’t understand (much more likely). After the completion of the Human Genome, the National Human Genome Research Institute (NHGRI) started a new program to map all of the functional DNA elements in the human genome. The goal was to figure out what the other 98% of the genome was doing by finding all of the regions and sequences in the human genome that contributed to human gene expression. This project is the basis of the ENCyclOpedia of Dna Elements (ENCODE) - one of the most awkward scientific acronyms ever conceived. They did this by performing a number of experiments to map gene expression, determine where proteins were binding to the genome, and determine how modifications to proteins control access to the DNA code.
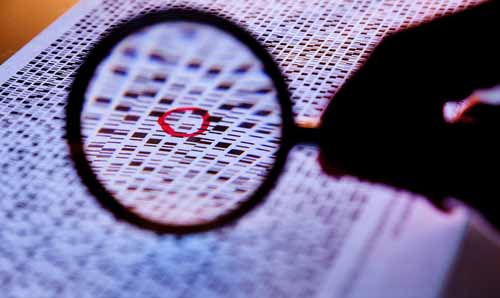
This week, the consortium published 30 papers explaining their results in aggregate. Unfortunately, the major theme that emerged from this extraordinarily important resource is that 80% of the genome does something if you define “does something” as “converted into RNA.” Headlines are ablaze with the “No Junk DNA” meme, but I’m going to let you in on a little secret. The scientific definition of the word junk is subtlely different than the definition that most of us assume. Most of us think that "junk" is defined as stuff that has no use, never will, and should be thrown out. Here, scientists have been defining junk for the past decade much more like a hoarder would. Junk DNA is the DNA that we currently have no known use for, but that may have a use in the future. Think of this like you would that "American Picker'" show on the History Channel where those guys drive around in a van to homes or properties filled with what much of us would consider worthless gharbage. Once in a while they pull out something cool or useful from that pile of "junk." So scientists are a lot like the "American Picker's", all along we thought that this DNA might have a use down the line because maintaining genomes is a costly business. Think of the genome as a computer hard drive. As your hard drive gets filled with data and the data becomes increasingly cluttered, your computer slows to a crawl and becomes impossible to use. At this point, you have one of two choices, you can trash your computer and buy a new one, or you can defragment the hard drive, get rid of the files you don’t use, and archive files that you think you might need. Evolution and our genome perform similar functions so it would make sense that the genome contains mostly useable information because at a certain point extraneous information is going to be too costly. One of the important stories from the ENCODE data is that by doing evolutionary comparisons of 24 mammalian genomes, the consortium was able to determine that negative selection of useless, neutral and negative elements is actively occurring. This tells us that the genome is taking care of the “junk,” but again, this isn’t even close to the most important information we can glean from the ENCODE datasets and it’s sad that so much of the media coverage has focused on the junk story line.
There’s a huge teachable moment here. Most of the public believes that DNA is what controls who we are and that the absolute sequence of the DNA is the most important bit of information. We know now that just isn’t the case, and yes, it is much more complex than that. Why and how proteins turn on specific genetic programs is dependent on much more than just the sequence of the DNA. Many genetic regulators bind to the genome in a sequence independent manner. Further, many of the start sites for genes expressed by the genome are not defined in a sequence specific manner. Every high school biology text talks about how the TATA box is the start site for DNA to be turned into RNA, except only 25% of the proteins expressed by the genome actually have a TATA site. For the majority of gene expression the genetic start sites are defined much more loosely as a function of which types of bases are present in the promoter or which proteins are bound there.
This is where the power of the ENCODE project comes in. We now know where a huge number of genetic regulators are located on the genome. This data underscores the fact that the DNA program executed in each cell type in your body is different! We know this from previous research, but we have a much better view of this today by looking at the ENCODE datasets. ENCODE has determined the genome binding location of a couple hundred very important gene expression regulators in 147 different cell types. Where these proteins bind, how they are modified and what stretches of DNA they have access to makes all the difference in determining whether a cell turns into a neuron, a muscle cell, or erupts into a life threatening melanoma. The scary thing is that the ENCODE data release only contains binding information for 119 of the 1,800 proteins we know are involved in regulating gene expression and the 147 cell types tested only lets us glance at the specific genetic recipes executed in the thousands of different cell types that make up our body. With so much information left to explore, the million dollar (trillion?) question becomes: Is all of this complexity knowable? Probably.
---
For more information and FREE ENCODE publications, visit the ENCODE portal at Nature: http://www.nature.com/encode
This post has been viewed: 11046 time(s)
 |
 |
Really if you use genetics as an analog to a computer the genome is like the hard drive on a computer of unknown architecture with an unknown file system running an operating system and programs. Currently we seem to be at the point where a disk image can be dumped and a few patterns can be recognised and fiddled with using a hex editor, then dumped back to the disk to see what the changes do.
Genetics is one of the many fields that could probably benefit from more cross pollination from other fields. We might even end up with at least a 2GL language for writing fresh genetic sequences from which more complex languages could be spawned. Understanding the pre-existing DNA and RNA may be an insurmountably massive task, but the ability to write genertic "patches" to help counter some conditions or add functionality would be invaluable.
There's quite a bit of research into creating DNA storage and DNA computers, but I was just using that as an easy to understand analogy about why having a lot of junk cluttering the genome could be a disadvantage. No doubt there's a ceratin amount of stuff we don't use anymore.
 |
 |
 |
 |









Jaeson, that's not true at most places. Top tier, sure, but 1100+ should get you past the first filter of most PhD programs in the sciences. . . .Read More
All I can say is that GRE's really do matter at the University of California....I had amazing grades, as well as a Master's degree with stellar grades, government scholarships, publication, confere. . .Read More
Hi Brian, I am certainly interested in both continuity and accuracy of PacBio sequencing. However, I no longer fear the 15% error rate like I first did, because we have more-or-less worked . . .Read More
Great stuff Jeremy! You bring up good points about gaps and bioinformatics. Despite the advances in technology, there is a lot of extra work that goes into assembling a de novo genome on the ba. . .Read More
Brian,I don't know why shatz doesn't appear to be concerned about the accuracy of Pacbio for plant applications. You would have to ask him. We operate in different spaces- shatz is concerned a. . .Read More